Command Palette
Search for a command to run...
THINGS-data: A multimodal collection of large-scale datasets for investigating object representations in human brain and behavior
THINGS-data: A multimodal collection of large-scale datasets for investigating object representations in human brain and behavior
Martin N Hebert Oliver Contier Lina Teichmann Adam H Rockter Charles Y Zheng Alexis Kidder Anna Corriveau Maryam Vaziri-Pashkam Chris I Baker
Abstract
Understanding object representations requires a broad, comprehensive sampling of the objects in our visual world with dense measurements of brain activity and behavior. Here, we present THINGS-data, a multimodal collection of large-scale neuroimaging and behavioral datasets in humans, comprising densely sampled functional MRI and magnetoencephalographic recordings, as well as 4.70 million similarity judgments in response to thousands of photographic images for up to 1,854 object concepts. THINGS-data is unique in its breadth of richly annotated objects, allowing for testing countless hypotheses at scale while assessing the reproducibility of previous findings. Beyond the unique insights promised by each individual dataset, the multimodality of THINGS-data allows combining datasets for a much broader view into object processing than previously possible. Our analyses demonstrate the high quality of the datasets and provide five examples of hypothesis-driven and data-driven applications. THINGS-data constitutes the core public release of the THINGS initiative (https://things-initiative.org) for bridging the gap between disciplines and the advancement of cognitive neuroscience.
One-sentence Summary
THINGS-data is a large-scale multimodal collection of neuroimaging and behavioral datasets designed to investigate object representations by integrating dense functional MRI and magnetoencephalographic recordings with 4.70 million similarity judgments across thousands of photographic images for up to 1,854 object concepts.
Key Contributions
- The paper introduces THINGS-data, a large-scale multimodal collection of neuroimaging and behavioral datasets designed to study object representations in humans.
- This dataset integrates densely sampled functional MRI and magnetoencephalographic recordings with 4.70 million similarity judgments across up to 1,854 object concepts.
- The study demonstrates the utility of this multimodal framework through five distinct hypothesis-driven and data-driven applications that showcase the high quality of the collected data.
Introduction
The provided text contains only acknowledgements, funding information, and author contributions, which do not describe the research background, technical context, or the specific scientific contributions of the study. Consequently, there is insufficient information to summarize the research objectives or the limitations of prior work.
Dataset
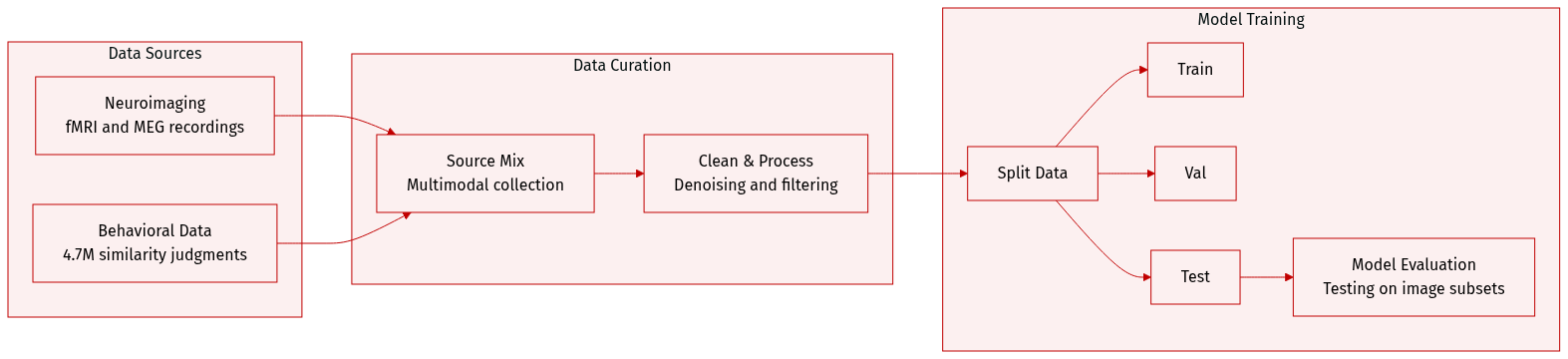
Dataset Description: THINGS-data
THINGS-data is a large-scale multimodal collection designed to investigate object representations in the human brain and behavior. The authors draw from the THINGS database, which contains 1,854 object concepts and 26,107 naturalistic images.
Dataset Composition and Sources The collection consists of three primary modalities:
- Functional MRI (fMRI): Neuroimaging data captured from 3 healthy participants.
- Magnetoencephalography (MEG): High-temporal-resolution neuroimaging data captured from 4 different healthy participants.
- Behavioral Data: Perceived similarity judgments collected via online crowdsourcing on Amazon Mechanical Turk from 12,340 workers.
Key Subset Details
- fMRI Subset: Includes 8,740 unique images representing 720 object concepts. To ensure data reliability, the authors used a subset of 100 images for noise ceiling estimation and model testing.
- MEG Subset: Includes 22,448 unique images representing all 1,854 object concepts. A subset of 200 images was used for noise ceiling estimation and model testing.
- Behavioral Subset: Comprises 4.70 million similarity judgments collected through a triplet odd-one-out task. This includes 37,000 within-subject triplets to estimate subject-level variability and additional subsets to assess data consistency.
Data Processing and Metadata
- Image Processing: Images were cropped to a square format, with padding applied to objects that did not fit. Stimuli subtended 10 degrees of visual angle against a mid-grey background.
- Neuroimaging Processing:
- fMRI data underwent slice timing correction, head motion correction, susceptibility distortion correction, and spatial alignment using fMRIPrep and Freesurfer.
- MEG data were bandpass filtered (0.1 to 40 Hz), epoch-based, and down-sampled to 200 Hz.
- The authors implemented a custom ICA-based denoising method for fMRI to separate signal from physiological noise.
- Eye-tracking: MEG sessions included 1,200 Hz eye-tracking to monitor central fixation. Data were processed to remove samples related to eyeblinks and pupil dilation spikes.
- Metadata Construction: The dataset is enriched with semantic and image annotations, including high-level categories, typicality, object feature ratings, and memorability scores.
Data Usage in the Paper The authors use the data to bridge spatial and temporal brain responses with human behavior. Specific applications include:
- Model Evaluation: Using the designated test image subsets to evaluate computational models.
- Representational Analysis: Performing multivariate pairwise decoding, encoding analyses (specifically for animacy and size), and representational similarity analysis (RSA).
- Multimodal Fusion: Implementing a regression-based approach to combine MEG and fMRI responses to uncover spatiotemporally resolved information flow.
Method
Single-Trial fMRI Response Estimation
To address the computational challenges and noise inherent in large-scale fMRI voxel-wise time series, the authors implement a method to estimate the BOLD response amplitude for each individual object image. This is achieved by fitting a single-trial general linear model (GLM) to the preprocessed time series. The procedure begins by converting the data from each functional run into percent signal change. To isolate the task-related signal, the authors regress the time series against a set of noise regressors, which include ICA noise components specific to each run and polynomial regressors up to the fourth degree. The residuals from this regression are retained for subsequent analyses.
To account for physiological variability in the hemodynamic response function (HRF), the authors utilize a library of 20 different HRFs. For every voxel, a separate on-off design matrix is generated for each of the 20 HRFs. The best fitting HRF per voxel is determined by identifying the model that yields the largest amount of explained variance. Because the regressors in a fast event-related design are highly correlated, the authors employ fractional ridge regression to prevent overfitting. The regularization parameter for each voxel is optimized to maximize the predictive performance of held-out data using a leave-one-session-out cross-validation approach. The hyperparameter space for the regularization parameter is sampled finely, ranging from 0.1 to 0.9 in steps of 0.1, and from 0.9 to 1.0 in steps of 0.01.
After determining the optimal HRF and fractional ridge parameter for each voxel, a single-trial model is fit to obtain the beta coefficients. Since ridge regression can introduce biases in the scale of these coefficients, the authors apply a linear rescaling by regressing the regularized coefficients against the unregularized coefficients, using the resulting predictions as the final single-trial response amplitudes. This process transforms the high-dimensional time series into a compact set of beta weights, where each voxel is represented by a single value per image.
Behavioral Embedding via SPoSE
The authors derive a low-dimensional embedding from triplet odd-one-out judgments using the SPoSE model. This computational framework is designed to identify sparse, non-negative, and interpretable dimensions that underlie similarity judgments. The process begins by splitting the triplet data into training and test sets using a 90–10 split. The embedding is initialized with 90 random dimensions within the range of 0 to 1.
For any given triplet of images, the model computes the dot product of the 90-dimensional embedding vectors for all three possible pairs. A softmax function is then applied to these products to produce three predicted choice probabilities. The loss function used to update the embedding weights consists of two components: the cross-entropy loss (the logarithm of the softmax function) and an L1 norm to induce sparsity. The trade-off between these two terms is controlled by a regularization parameter λ, which is identified through cross-validation on the training set.
Optimization is performed using the Adam optimizer with a minibatch size of 100 triplets. Following optimization, dimensions with weights below 0.1 are removed, and the remaining dimensions are sorted in descending order based on the sum of their weights. To ensure the reproducibility of the stochastic optimization process, the authors run the procedure 72 times with different random seeds. They calculate a reproducibility index for each dimension based on the average Fisher-z transformed Pearson correlation across the different embeddings. The final embedding, which consists of 66 dimensions, is selected based on having the highest mean reproducibility.
Experiment
The researchers evaluated a large-scale behavioral, fMRI, and MEG dataset to validate its utility for studying object representations. By expanding the behavioral dataset to 4.7 million trials, they successfully captured more fine-grained similarity dimensions and improved predictions for within-category ratings. Neuroimaging analyses confirmed that both object identities and categories are reliably decodable, while replication studies demonstrated that established cortical gradients for animacy and size generalize to a much broader range of objects. Finally, integrating these modalities through representational similarity analysis and direct regression revealed that human similarity judgments are closely linked to both the spatial topography and temporal dynamics of neural responses.