Command Palette
Search for a command to run...
Cornell University Has Developed EMSeek, a multi-agent Platform That Can Transform Electron Microscope Images Into Materials Science Insights in Just 2-5 minutes.

Electron microscopy (EM) has provided humanity with an unprecedented window into the atomic world, allowing direct observation of defects, lattice distortions, and chemical inhomogeneities that determine the performance of catalysts, batteries, and semiconductors. While the volume of electron microscopy data has exploded, a significant problem remains that most datasets are still not fully analyzed:This is not because it lacks scientific value, but because the expert interpretation process is slow, fragmented, and difficult to reproduce.
In recent years, advancements in AI technology have offered new possibilities for unlocking the full scientific value of electron microscopy data. However, due to the diversity of imaging modalities and analytical tasks, existing applications remain largely confined to isolated steps. For example, AtomAI provides a pixel-level method for atomic-level segmentation, while AutoMat advances the automation of structure indexing.Each only covers a specific part of the EM analysis process.This fragmentation makes it difficult to connect the original microscopic images with crystallographic models, property predictions, or documentary evidence, thus failing to bridge the "last mile" from observation to insight.
Therefore, a key question arises:Is it possible to build a "virtual electron microscientist" capable of autonomously handling diverse imaging tasks, traversing different materials subfields, and integrating interdisciplinary knowledge?If an intelligent agent can manage tens of thousands of microscopic images simultaneously, it will greatly improve the efficiency of human scientific research and accelerate materials innovation.
In this context,A research team from Cornell University has proposed EMSeek, a modular multi-agent platform with source tracing capabilities.EMSeek integrates perception, structure reconstruction, property inference, and literature reasoning into a unified electron microscopy analysis workflow. Evaluation results on 20 material systems and 5 task categories show that EMSeek achieves approximately twice the speed and higher accuracy of Segment Anything in segmentation tasks; achieves structural similarity exceeding 90% on the STEM2Mat dataset; and, with calibration using only approximately 2% of labeled data, meets or exceeds the performance of strong single-expert models on three out-of-distribution property prediction benchmarks. More importantly,This method takes only 2 to 5 minutes to query each image completely, which is about 50 times faster than an expert process.
The related research findings, titled "Bridging electron microscopy and materials analysis with an autonomous agentic platform," have been published in Science Advances.
Research highlights:
* This research proposes a modular, multi-agent system with source tracing capabilities, unifying perception, structural modeling, property inference, and literature reasoning into a reproducible EM workflow.
* Case studies on two-dimensional lattices and supported nanoparticles validate EMSeek's generalization ability, demonstrating its potential to accelerate materials discovery and assist both professional and non-professional researchers.
EMSeek enables "virtual scientists" to collaborate with human researchers, accelerating materials discovery across the board, from fundamental characterization to device optimization.

Paper address:
https://www.science.org/doi/10.1126/sciadv.aed0583
Follow our official WeChat account and reply "EMSeek" in the background to get the full PDF.
Dataset: Balancing the breadth and difficulty of current electron microscopy analysis
To train and evaluate SegMentor, the core unit of this multi-agent system, researchers constructed a benchmark dataset that reflects the breadth and difficulty of current electron microscopy analysis, as shown in the figure below:
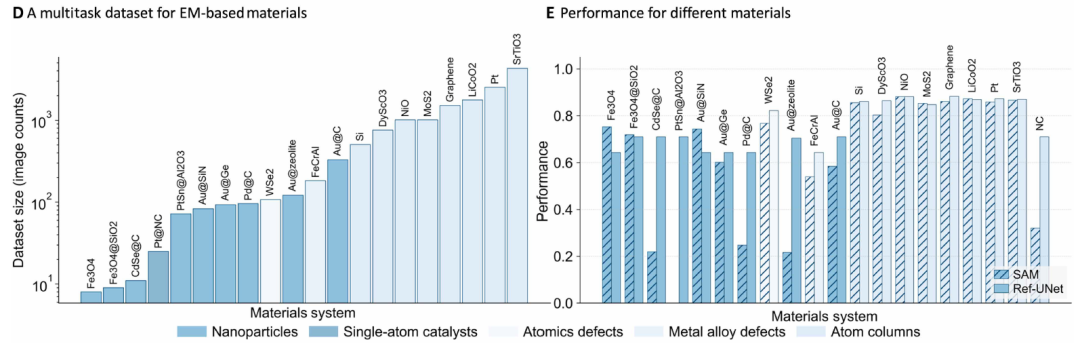
It covers 20 material systems and 5 mission types.
This dataset contains thousands of pixel-level annotated micrographs from 20 material systems (perovskites, high-entropy alloys, van der Waals heterostructures, single-atom catalysts, etc.).The tasks are divided into five categories: atomic column localization, point defect annotation, nanoparticle contour extraction, irradiation-induced defect counting, and single-atom identification.
Material selection follows three criteria:
(i) Related to key issues such as catalysis, energy storage, and semiconductor reliability;
(ii) It encompasses a wide range of structural complexities, from highly symmetric crystals to highly defective or low-contrast lattices;
(iii) It possesses imaging diversity, including variations in acceleration voltage, dose, and detection mode, which rigorously tests the robustness of the model.
More detailed statistical information is shown in the table below:
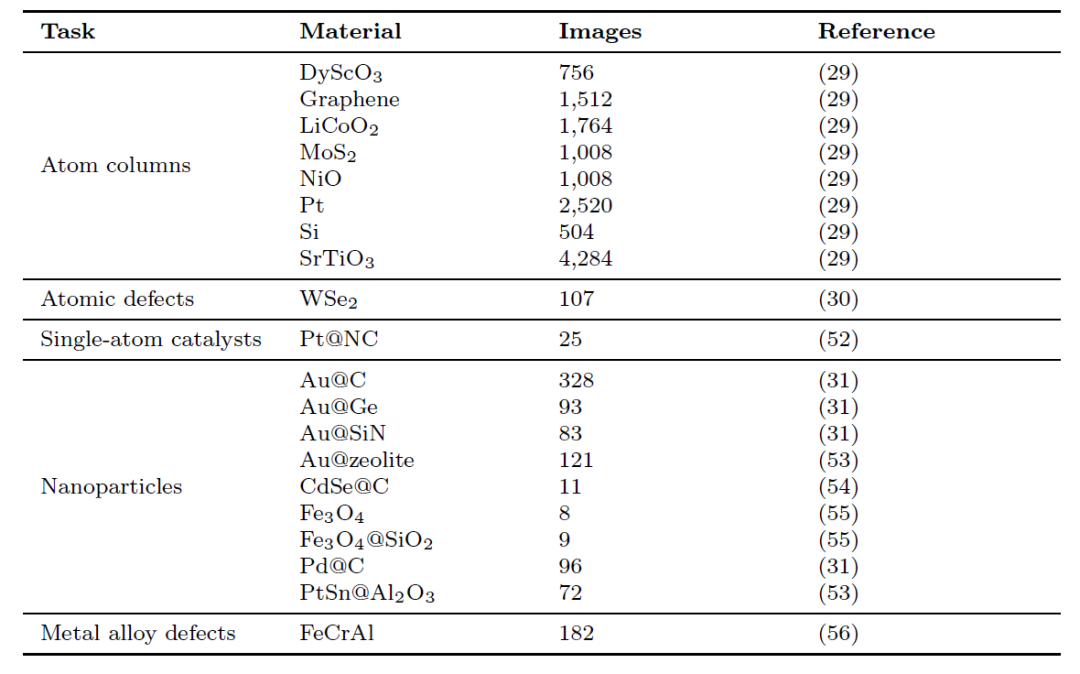
Public datasets are accompanied by references, while "Private" entries indicate data that has been annotated and compiled by the authors themselves.
EMSeek framework: Tasks are uniformly scheduled by a large language model.
Unlike relying on a single deep learning system,EMSeek assigns tasks to hierarchical specialized agents, which are then uniformly scheduled by Large Language Models (LLMs) to automatically complete planning, invocation, and execution.This minimizes human intervention, and its overall framework is shown in the figure below:
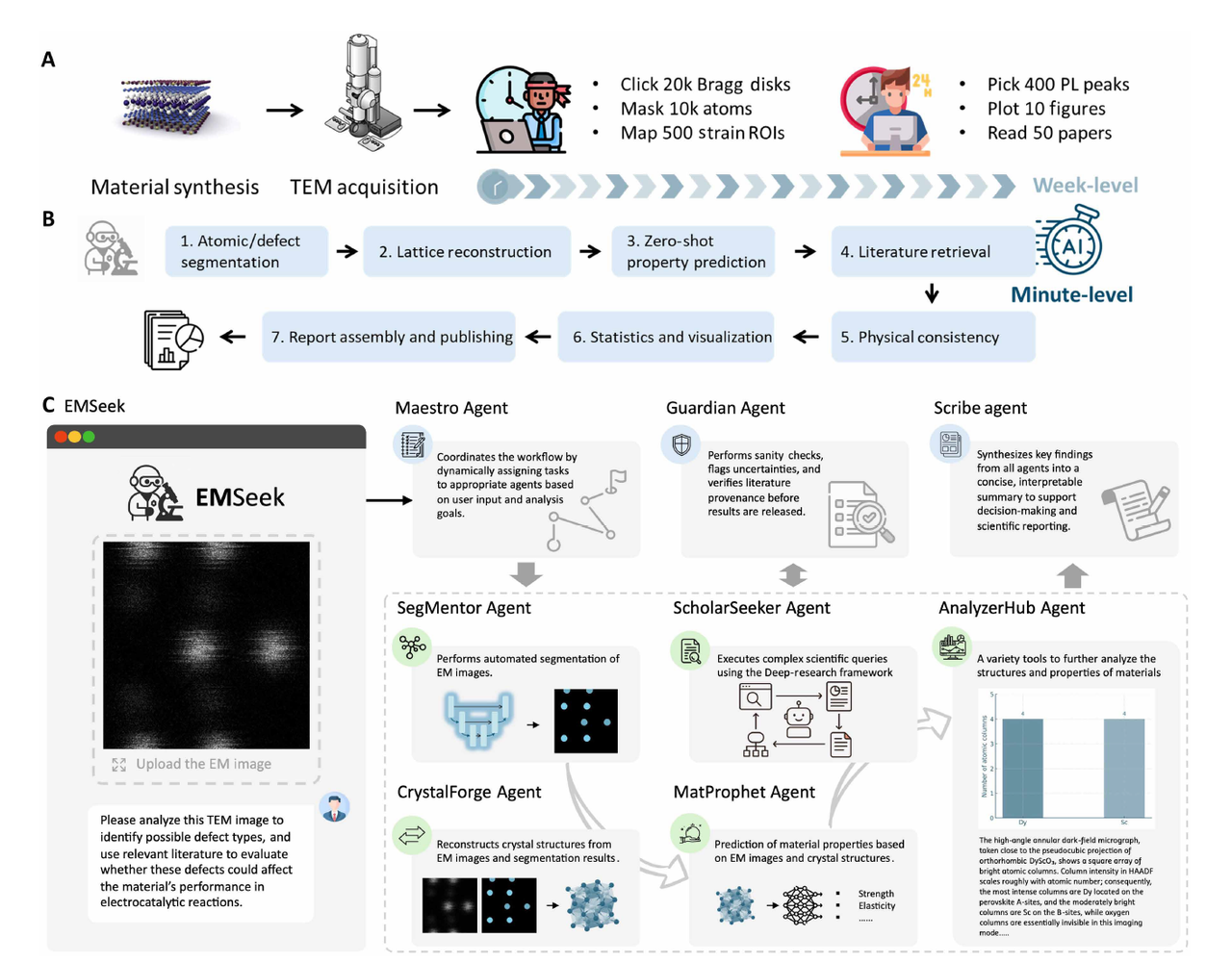
The platform comprises 5 core units, including:
SegMentor
This module is responsible for reference-guided "generic segmentation".Atomic-level and particle-level masks can be generated under various materials and imaging conditions. At the heart of SegMentor is Ref-UNet, a lightweight U-Net in which the encoder block is replaced by a visual backbone, and skip connections not only pass feature maps but also learned embeddings of user-selected reference patches.
During forward propagation, reference patches are tokenized, positionally encoded, and injected into each encoder stage via a cross-attention layer that reweights channel responses based on patch similarity. The generated context vectors propagate along the upsampling path, guiding pixel-level predictions to approach features that match the reference while suppressing interference.
CrystalForge (EM2CIF)
This module performs a reciprocal space search under mask constraints and combines database retrieval with candidate generation to reconstruct unit cell structures that can be used for density functional theory (DFT), even when faced with unknown chemical systems.
MatProphet
This module employs a gated expert hybrid (MoE) model, fusing the outputs of multiple inter-atom models.Calibration can be completed with only about 2% of labeled data.It also predicts properties such as formation energy and defect energy, as well as their uncertainties;
ScholarSeeker
This module is responsible for retrieving and synthesizing evidence from a massive amount of literature to generate answers with citation anchoring, thereby reducing the "illusion."It operates in a three-stage loop: Document retrieval: Retrieve candidate paragraphs through dense similarity queries; Evidence extraction: Sort and filter sentences, and record the complete source (DOI and sentence offset) through the Guardian Agent; Reasoning analysis: Organize the evidence into a structured argument, and inject the cited and supporting text into the user report by the Scribe Agent.
Guardian
At each handover step, physical rationality, unit consistency, and traceability information are verified, and Scribe integrates the mask, crystal structure file (CIF), property data sheet, and references into an auditable report.
EMSeek platform simplifies materials research
The EMSeek platform simplifies materials research by identifying key features in microscopic images, determining crystal structures, predicting material properties, comparing results with existing scientific literature, and generating reports within an integrated workflow. Researchers have validated EMSeek's comprehensive capabilities through a series of experiments.
EMSeek achieves universal atomic segmentation through "one-click guidance".
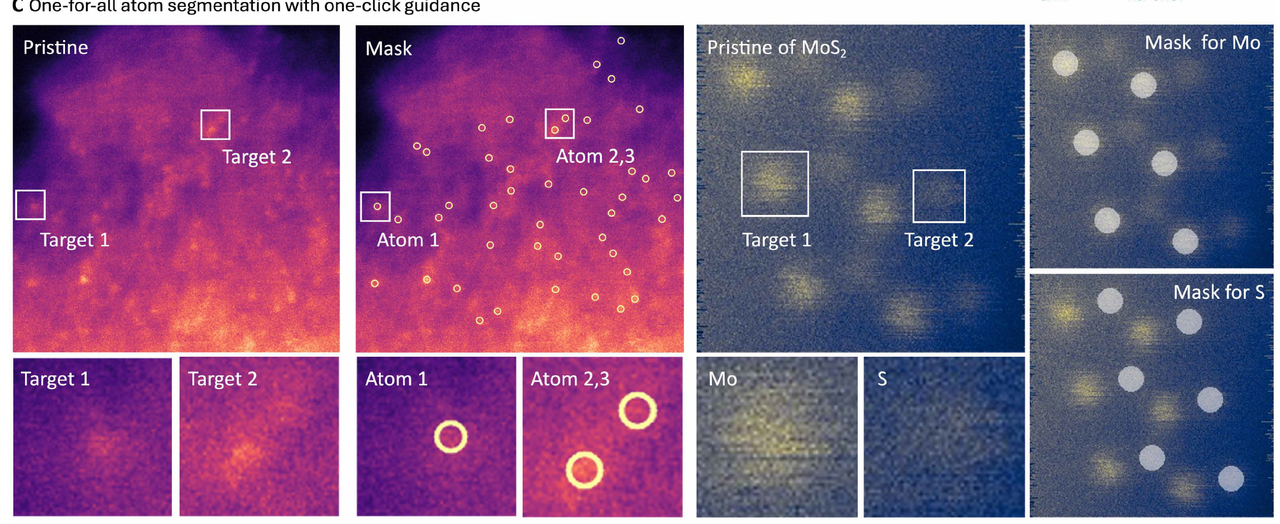
High-resolution electron micrographs typically contain tens of thousands of atomic columns in a single micrograph, but it is difficult to identify all crystallographically equivalent sites in a single processing run, whether by manual methods or cue-based tools. EMSeek proposes a "one-for-all" atomic segmentation mode: users only need to click on a representative site, and the model can automatically search and label all crystallographically similar atomic columns across the entire field of view.
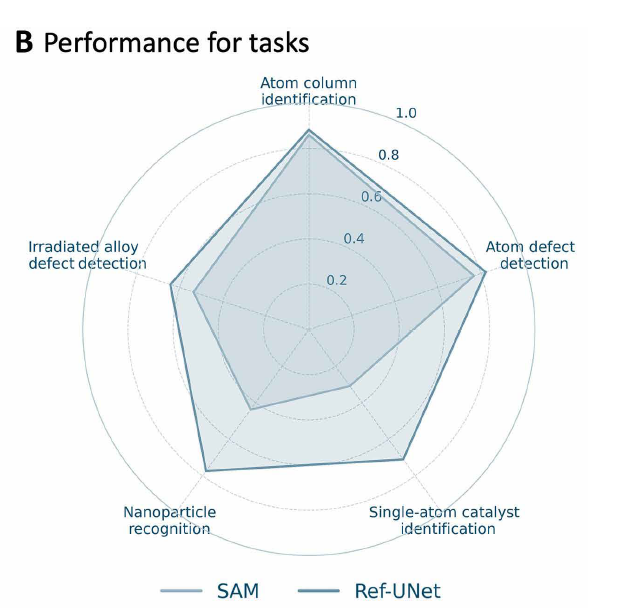
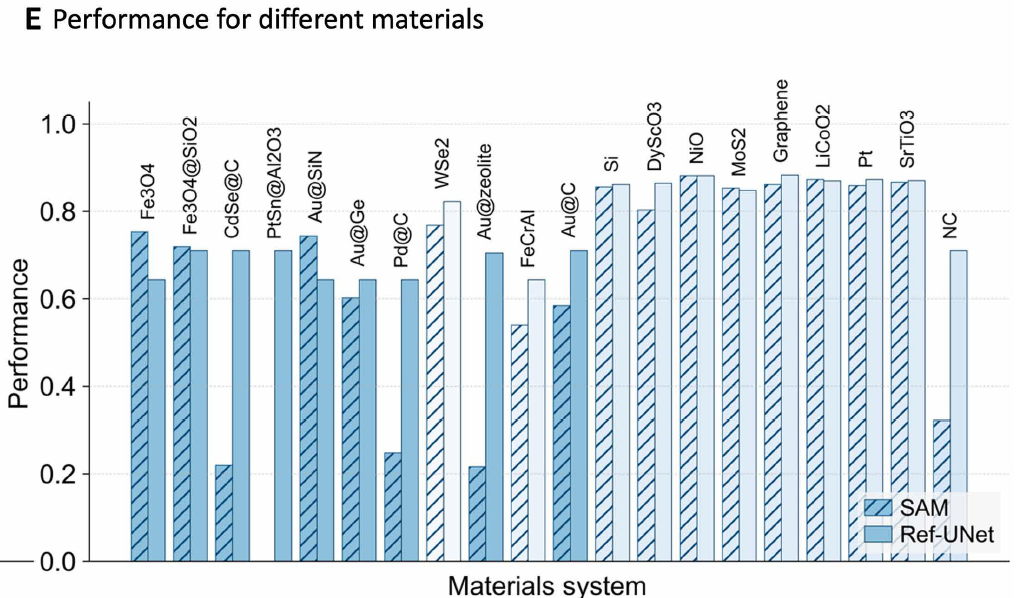
Researchers trained Ref-UNet using a multi-task dataset. Experiments show that Ref-UNet significantly outperforms the Segment Anything Model in both task performance and overall performance (see Figures B and E below). Its computational cost is 259 GFLOPs, and its parameter count is 28 million, only about half that of SAM 2 Hiera-B+ (560 GFLOPs, 81 million parameters).It achieves approximately twice the inference speed on a single GPU, thereby supporting real-time interactive feedback in the EMSeek workflow.
In terms of system implementation, researchers encapsulated Ref-UNet as a segmentation agent for EMSeek and combined it with an intuitive electron microscope image viewer to achieve real-time inference. As shown in the figure below,Each user click is converted into a reference tensor input model and returned as a superimposed result within a single frame, enabling researchers to iteratively optimize the segmentation mask while browsing tilt sequences or in-situ videos.The agent can also export pixel-level accurate regions of interest (ROI), statistical descriptions, and three-dimensional atomic coordinates, which can be directly input into the automatic CIF construction, phase fraction analysis, and subsequent property prediction modules.

EMSeek bridges the gap between electron microscopy and crystallography with its "one-click CIF generation" feature.
Electron microscopy images are crucial for resolving atomic structures, but converting noisy two-dimensional projections into reliable crystallographic models is often a fragile and labor-intensive process. Traditional methods typically rely on a combination of global template matching and manual indexing, but are extremely sensitive to factors such as contrast drift, stage or scanning instability, carbon contamination, and partial occlusion by the Bragg disk.
EMSeek avoids these problems by using a "segment first, then rebuild" strategy.Compared to pixel-based methods, EMSeek outperforms real-world structures in terms of structural similarity: its structural similarity exceeds 90% across all three difficulty levels, and it surpasses AtomAI and AutoMat at all levels (Figure C, left below). This advantage is particularly pronounced in high-noise scenes, as minute changes in contrast or residual carbon film can often cause significant shifts in global matching methods.
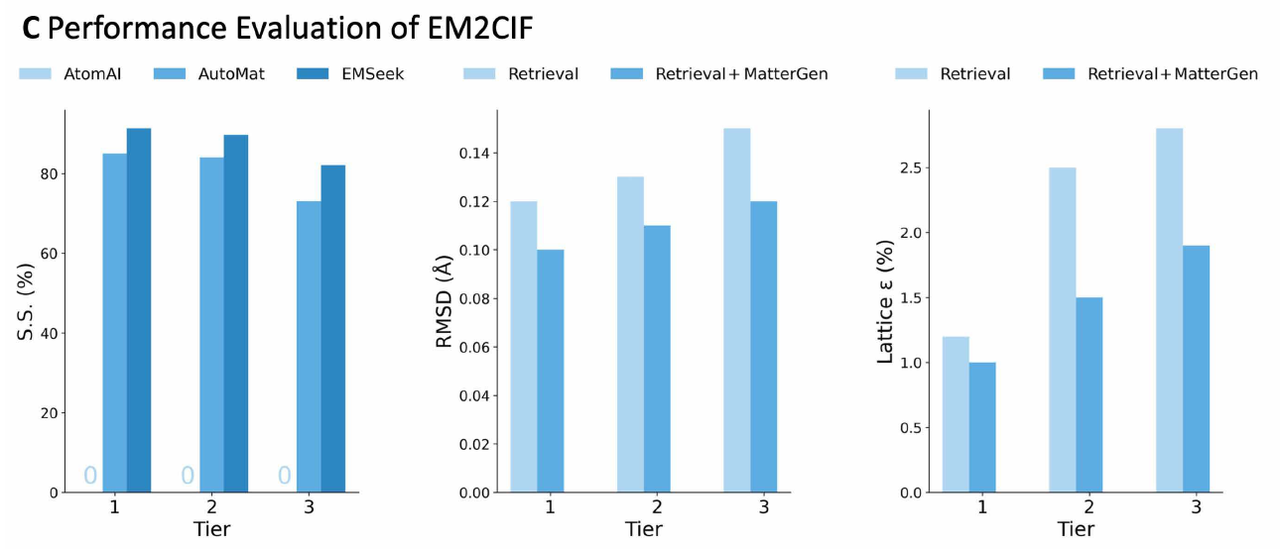
Benchmark tests were conducted on structural similarity (SS), root mean square deviation (RMSD), and lattice error (ε) at different difficulty levels, and EMSeek was compared with AtomAI, AutoMat, and pure search and hybrid modes.
In addition to its robustness, EMSeek is also able to adapt to material systems that go beyond the scope of existing databases.Therefore, it is not only suitable for stable structure reconstruction under normal conditions, but also provides reliable support in exploratory scenarios with high noise, polymorphic imaging conditions and unknown lattice structures.
EMSeek's generalization ability in multi-domain real-world materials tasks
Materials problems based on electron microscopy data are highly diverse. Traditional automation methods typically rely on custom scripts, temporary data formats, and extensive manual processing. Issues such as low signal-to-noise ratio, overlapping defect and strain comparisons, electron beam drift, and scarce annotations further hinder the realization of fully automated workflows. Even with frameworks like AtomAI, analysts often need to retrain models for each new dataset, taking weeks. EMSeek breaks this fragmentation by translating user requests into natural language prompts and integrating them into a unified intelligent agent framework.
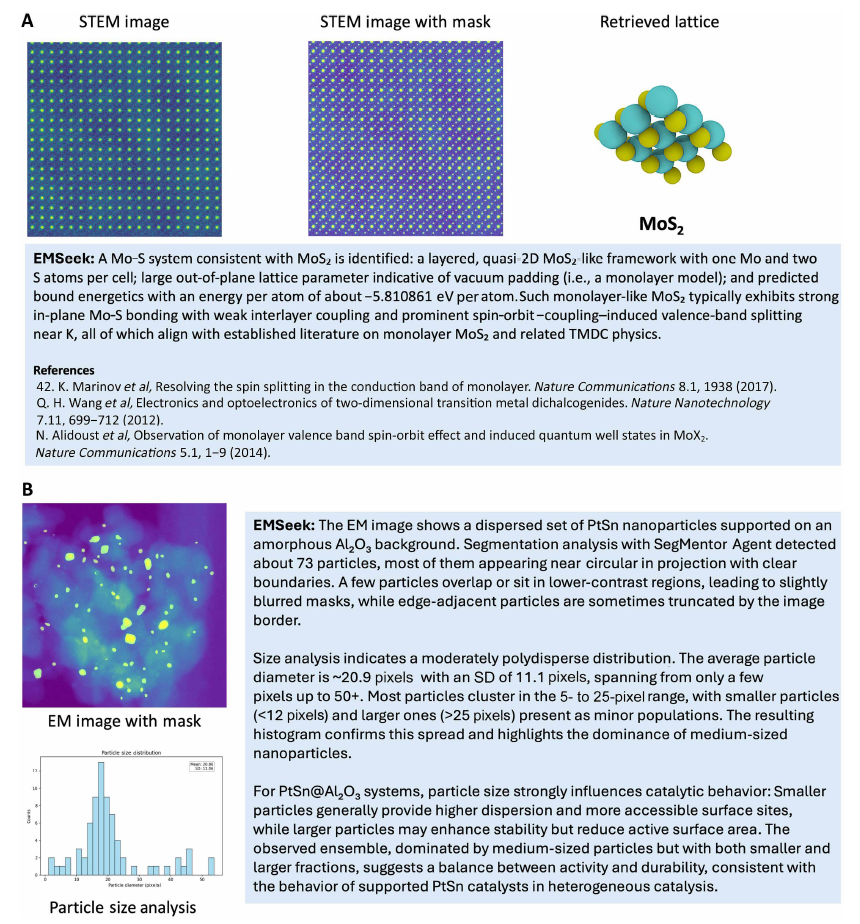
Figure A below shows the lattice identification of a monolayer-like MoS₂ in a scanning transmission electron microscope (STEM) image: SegMentor performs Mo and S atomic column segmentation, the mask is passed to CrystalForge (EM2CIF) to reconstruct the MoS₂ lattice, and then MatProphet predicts material properties (such as energy per atom). Figure B below shows the analysis of PtSn nanoparticles loaded on amorphous Al₂O₃: SegMentor detects approximately 73 approximately spherical particles, and AnalyzerHub generates a size distribution histogram showing moderate polydispersity. All mask, particle statistics, and histograms are recorded and traceable. The entire process takes only a few seconds per micrograph, and all intermediate results are traceable.
In addition to image analysis,EMSeek also integrates document-based reasoning capabilities for scientific question answering.As shown in the figure below, after incorporating domain literature through ScholarSeeker, the accuracy and verifiability of responses on the Metallurgy-QA dataset are superior to basic large language models or simple web searches. It is worth noting that simple network augmentation can sometimes actually reduce performance because large-scale web retrieval results often contain information that is weakly relevant to the task, thus introducing evidence mismatch or method substitution problems.
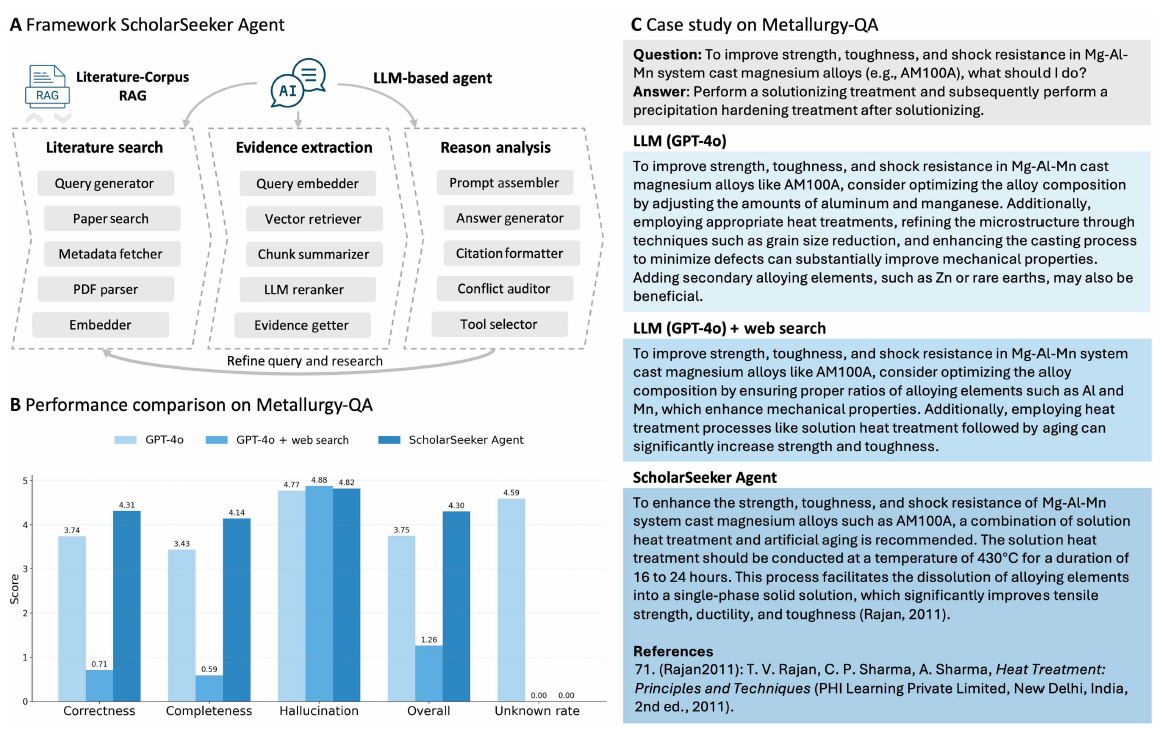

In contrast, by retrieving and extracting evidence from peer-reviewed literature, the system can significantly reduce "illusions" and improve the accuracy and completeness of answers. For example, when answering the question of how to improve the strength and toughness of Mg-Al-Mn alloys, the system can retrieve authoritative heat treatment processes and provide precise recommendations with citations.
EMSeek compresses electron microscopy analysis from weeks to minutes.
In a recent in-situ transmission electron microscopy (TEM) study,It took three experts nearly 20 weeks to complete the comprehensive annotation of 1200 frames of irradiation defects.Conventional static images typically require several minutes to an hour per image to reliably count defects. Atomic resolution lattice analysis or grain mapping also requires hours of expert work and is susceptible to analyst bias.
On the same type of data, EMSeek can complete reference-guided segmentation, mask-aware lattice reconstruction, MatProphet MoE property prediction, literature search and verification, and report generation on four A100 GPUs.It only takes 146 ± 18 seconds.This processing speed reduces actual turnaround time by two to three orders of magnitude, transforming electron microscopy from a post-hoc diagnostic tool into a near real-time hypothesis testing and process optimization driver.
New competitive focus in future materials research
From a broader perspective, artificial intelligence is reshaping the fundamental paradigm of materials science research, particularly unleashing unprecedented efficiency in the critical link of "characterization-understanding-design." Traditionally, atomic-level materials characterization has heavily relied on experienced experts, requiring months or even years of professional training. Even seasoned operators struggle to guarantee the stability and consistency of results when dealing with novel systems such as two-dimensional (2D) materials. This strong dependence on human resources and experience has long been a bottleneck for the scaling and automation of materials research, directly driving the development of intelligent characterization systems that require "less data and have lower barriers to entry."
A representative example is the ATOMIC (Autonomous Technology for Optical Microscopy and Intelligent Characterization) framework proposed in 2025 by Wang Haozhe of Duke University and Ren Zhichu's team at MIT.This is an end-to-end framework integrating a base model to achieve fully autonomous zero-sample characterization of 2D materials. The system integrates a visual base model (i.e., the Segment Anything model), a large language model (i.e., ChatGPT), unsupervised clustering, and topological analysis, automating microscope control, sample scanning, image segmentation, and intelligent analysis through rapid engineering without additional training. In analyzing typical MoS(2) samples, the method achieved a single-layer recognition and segmentation accuracy of 99.71 TP3T, comparable to that of professionals.
Paper title: Zero-shot Autonomous Microscopy for Scalable and Intelligent Characterization of 2D Materials
Paper link:
https://arxiv.org/abs/2504.10281
Echoing EMSeek's multi-agent framework, ATOMIC demonstrates another path: transforming complex experimental processes into programmable and reusable intelligent tasks through end-to-end automation driven by a base model. The common value of these systems lies in the fact that they no longer rely on large-scale labeled data or repeated training for a single task, but rather achieve rapid adaptation to new material systems through a combination of general capabilities.This means that one of the key areas of competition in future materials research will shift to "who can more efficiently manage intelligent agents and knowledge resources".
It is foreseeable that materials science will gradually enter an era of "AI-native" research: experimental data can be analyzed instantly, model predictions and literature knowledge can be linked in real time, and researchers will be freed from heavy data processing and repetitive work, allowing them to focus on formulating questions and designing experiments. In this process, artificial intelligence is no longer just an auxiliary tool, but is becoming a core infrastructure driving materials discovery and innovation.
References:
https://www.science.org/doi/10.1126/sciadv.aed0583
https://phys.org/news/2026-04-ai-electron-microscopy-materials-insights.html
https://mp.weixin.qq.com/s/AaAHOpChVXj_2xQJRvqHSg